What is KMeans Clustering?

KMeans Clustering is a clustering algorithm, which is to say it takes points of data, and tries to discern groups, or clusters, of those points based on their distance from k number of centroids. These centroids are originally chosen as random points within the scope of the data, then slowly altered by finding the mean of the nearest points to each centroid. Once the groups of points linked to each centroid stops moving, the model has “converged” meaning that further iteration wouldn’t change anything.
An Example, the UCI Iris dataset:
Lets say you’re given these two graphs and told to find how many classes are in the data based on them:
| Petal Width | Petal Length |
|---|---|
 |
 |
At first glance, it’s not that clear what the data is saying. It’s obvious that there’s two distinct groups in both the width and length values, but one portion seems to be a larger, wider blob that’s more spread out. Let’s try adding these two dimensions together.
A starter solution would be to plot the points as a scatter plot:

Now the data is a bit more visible, and it’s obvious that there are two distinct groups. While we’re still not displaying all dimensions of the data (the Iris dataset has four variables to work with), we can still use a clustering algorithm like KMeans to cluster the data using all four dimensions. Also, while it is already known from the data set that there are three classes in the data, lets say for the sake of experimentation that we don’t know and want to use KMeans to find out.
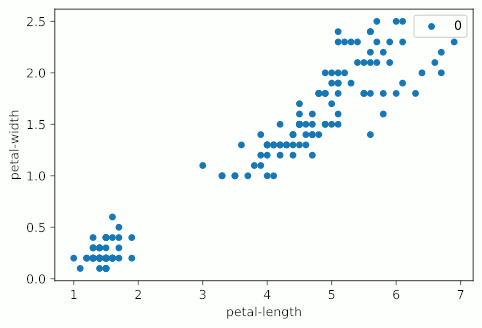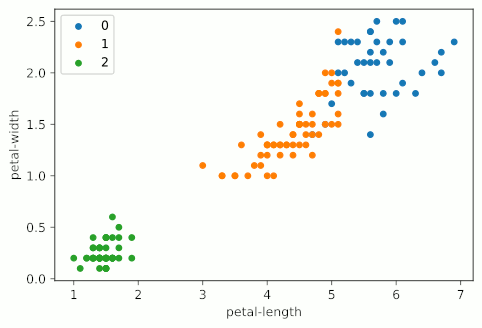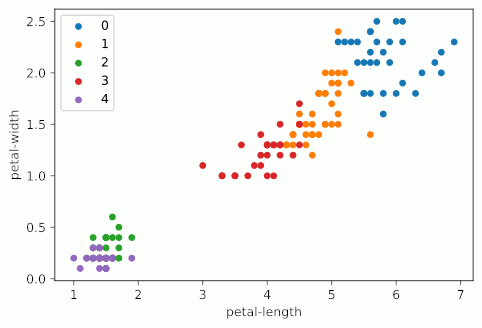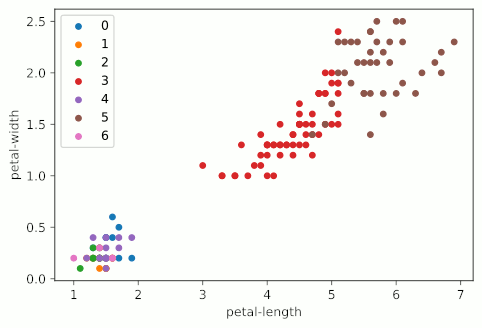
It looks like KMeans starts making stuff up around four clusters, and really starts falling apart at five clusters, when the bottom left group gets shattered into a bunch of colors. It should also be noted that my graphs are not as uniform as the example graph at the top of the page. This is primarily due to the fact that I’m using all four variables of the Iris dataset, but only displaying two; I’m essentially taking a 2D slice of a 4D graph.
How it works
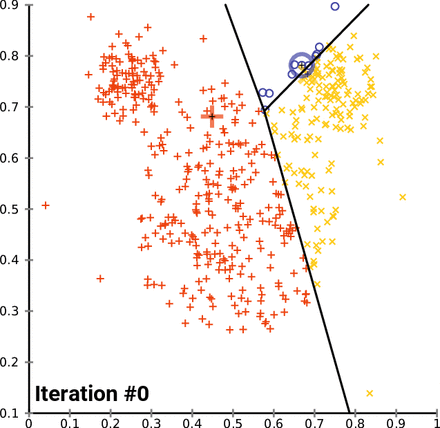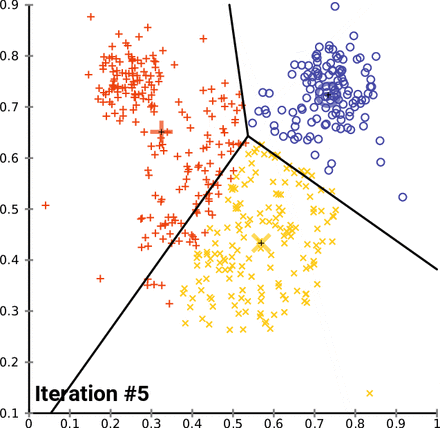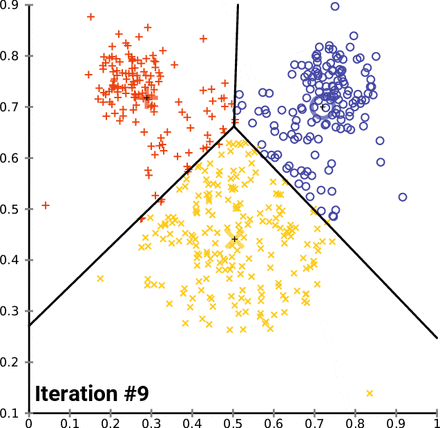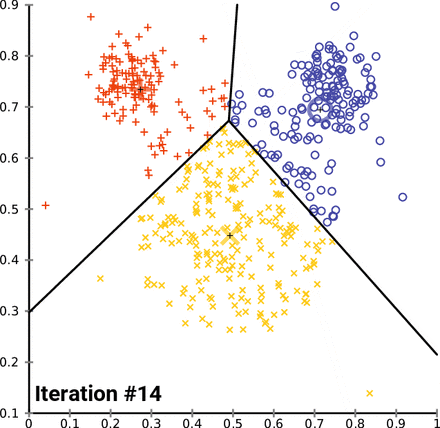
KMeans starts by picking k random points, called centroids, within the scope of the dataset, where k is the number of clusters you want KMeans to look for. From there, it groups the points of the dataset by which centroid it is nearest to. From there, it gets the mean point of each group - essentially the average, and designates those points as the new centroids, and then runs again. KMeans will do this as many times as it needs to until the model “converges”, which is to say that the centroids don’t change when the model tries to update them. In the case of the Iris dataset, it looks something like this:
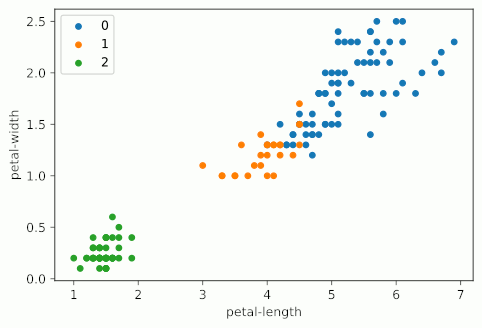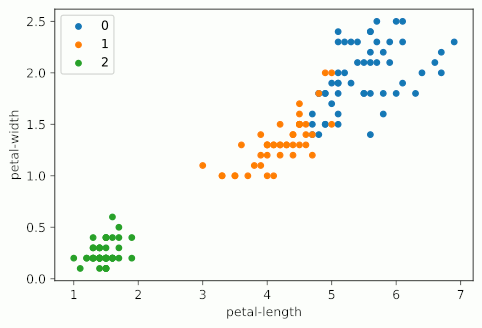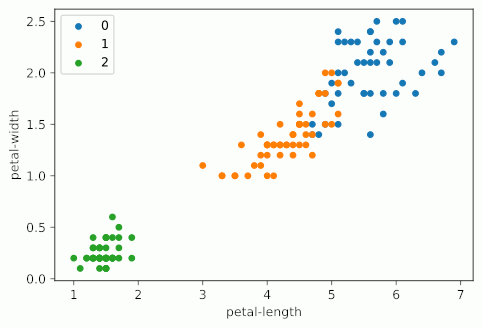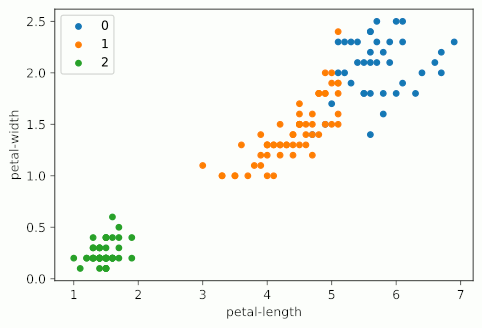
Notice how as KMeans iterates, cluster 1 slowly creeps up cluster 0 as the centroids for each are updated; cluster 2 probably doesn’t change at all due to its remoteness from the rest of the data, but maybe some points shift on the two dimensions not shown on the graph.
How it works (my code)
KMeans has two essential functions fit() and predict(). fit() is the function that sets up k number of centroids to fit with the data set given, and predict() is the function that takes the data set and returns a list of labels matching those data points with what cluster they ended up in. Outside of the math module built into Python, which I use for math.sqrt(), there are no libraries that need to be installed to run my code.
fit()
fit() starts by initializing k centroids, which for my code is based on a variable set during initialization called n_clusters. While standard KMeans procedures call for setting the initial centroids to be random points within the data’s scope of values for each dimension, my code just grabs the first k number of points and uses those:
sample_idx = [x for x in range(self.n_clusters)]
self.centroids_ = [data[x] for x in sample_idx]
From there, it enters the main loop where the iteration to converge the model takes place:
iter_count = 0
# loop through iterating centroids; stop if max iterations reached
while iter_count < self.max_iter:
prev_labels = self.labels_
# reset labels and calculate new ones
self.labels_ = []
for row in data:
dists = []
for c in self.centroids_:
dist = float(0)
# for each dimension, get the absolute value
# of the distance and calculate total distance
# via pythagorian theorem
for i in range(len(row)):
dist += abs(row[i]-c[i]) ** 2
dist = math.sqrt(dist)
dists.append(dist)
# go through each distance and get the smallest
for i in range(self.n_clusters):
if dists[i] == min(dists):
self.labels_.append(i)
# stop early if the model's converged
if self.labels_ == prev_labels:
break
# calculate new centroids
pointlist = [[] for x in self.centroids_]
self.centroids_ = []
for i in range(len(data)):
pointlist[self.labels_[i]].append(data[i])
for k in range(self.n_clusters):
means = [0 for x in range(len(pointlist[k][0]))]
# add up distances for each dim
for row in pointlist[k]:
for n in range(len(row)):
means[n] += row[n]
# divide by length of pointlist for each dim
for n in range(len(pointlist[k][0])):
means[n] = means[n] / len(pointlist[k])
# append new centroid
self.centroids_.append(means)
iter_count += 1
predict()
predict() is essentially just a repeat of the second half of fit(), generating a list of labels based on the centroids stored in KMeans that were created by fit()
# initialize list of labels to be returned
labels = []
for row in data:
dists = []
for c in self.centroids_:
dist = float(0)
# get absolute value of distance
# on each dimension
for i in range(len(row)):
dist += abs(row[i]-c[i]) ** 2
dist = math.sqrt(dist)
dists.append(dist)
# go through each distance and get the smallest
for i in range(self.n_clusters):
if dists[i] == min(dists):
labels.append(i)
return labels
How it works (compared to sklearn.cluster.KMeans)
Obviously, this problem has been solved before; writing a KMeans algorithm is nothing new and there are probably loads of implementations of the algorithm, the most popular of which, for Python, is Sci-Kit Learn’s implementation of KMeans. Comparing my code to SKLearn’s wouldn’t do it any justice, as the point of me going out of my way to write a KMeans algorithm from scratch was to show how it works and how I went about doing so. A better test of my code against SKLearn’s would be to compare two graphs of the same data, one run through each algoritm, to show the difference in results.
| My Implementation | SKLearn’s Implementation |
|---|---|
 |
 |
Aside from the different label colors which probably arose from SKLearn’s algorithm starting with random points, as well as one dot being a different color, our graphs are exactly the same, meaning my implementation is spot-on with what KMeans is designed to do.
Resources
SKLearn KMeans implementation: link
My KMeans implementation: link
My KMeans source code: link
SKLearn’s KMeans doc: link
SKLearn’s KMeans source code: link